Deploy ML Models: Real-Time Alerts & Batch Screening
Level: Intermediate
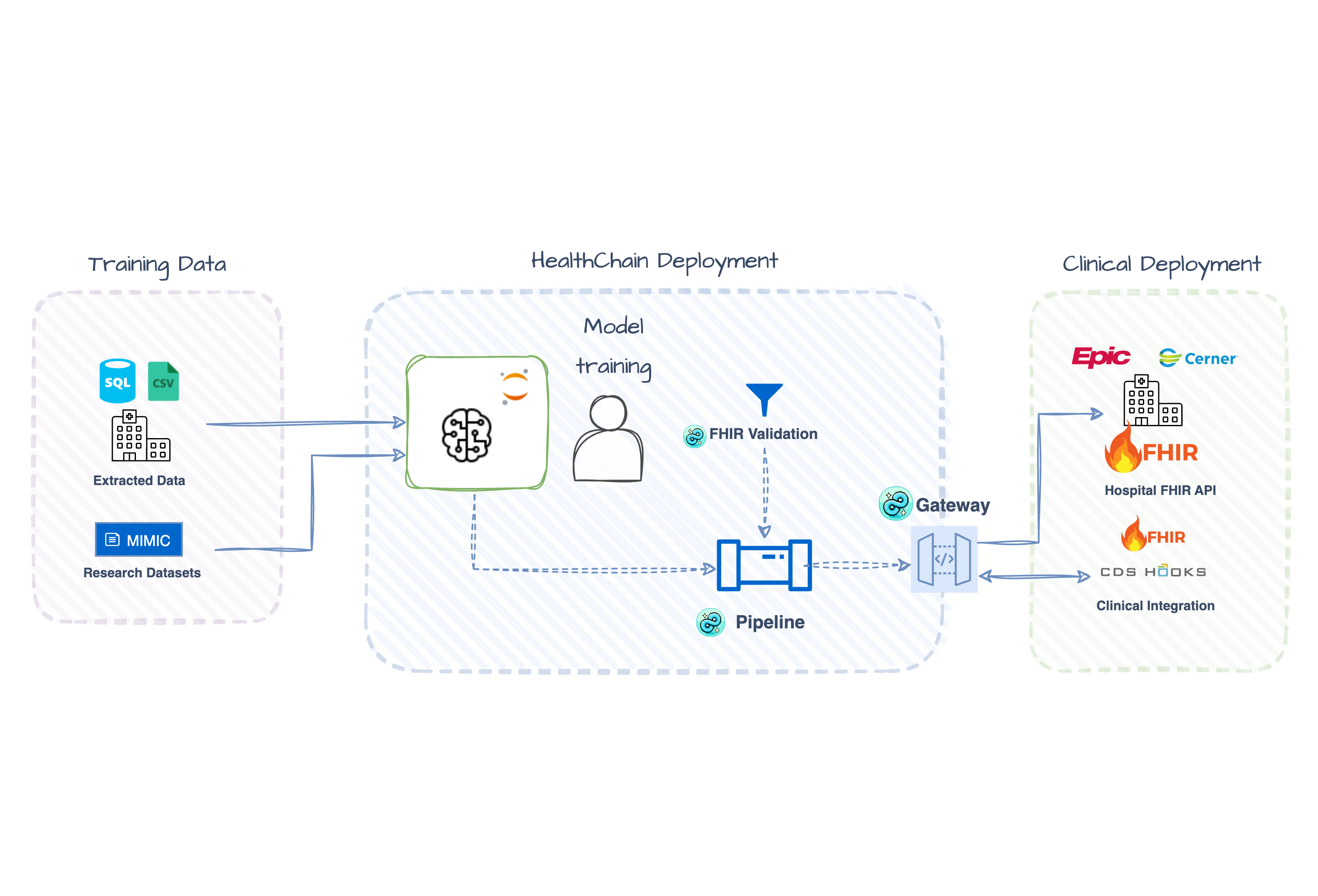
You trained a model on CSVs. Now you need to deploy it against FHIR data from EHRs. This tutorial shows how to bridge that gap with two production patterns: real-time CDS Hooks alerts and batch FHIR Gateway screening — both using the same model and a simple YAML schema that maps FHIR resources to your training features.
Check out the full working examples:
When to Use Each Pattern
| Pattern | Trigger | Output | Best For |
|---|---|---|---|
| CDS Hooks | Clinician opens chart | Alert cards in EHR UI | Point-of-care decision support |
| FHIR Gateway | Scheduled job / API call | RiskAssessment resources | Population screening, quality measures |
Both patterns share the same trained model and feature extraction — only the integration layer differs.
Quick Start: CDS Hooks in 5 Minutes
The demo patients and a pre-generated model are already in the repo — no training or data download needed.
That's it. The script starts a local CDS Hooks service, fires test requests against it using three pre-extracted MIMIC patients, and prints risk scores:
Processed 3 requests
Patient 1: Sepsis Risk: HIGH (85%)
Patient 2: Sepsis Risk: MODERATE (52%)
Patient 3: Low risk (no alert)
Results are saved to ./output/. The rest of this tutorial explains how it works and how to adapt it.
The Shared Model Pipeline
Both patterns reuse the same pipeline. It loads a pre-trained XGBoost classifier and runs inference on a Dataset extracted from a FHIR Bundle:
def create_pipeline() -> Pipeline[Dataset]:
pipeline = Pipeline[Dataset]()
@pipeline.add_node
def impute_missing(dataset: Dataset) -> Dataset:
dataset.data = dataset.data.fillna(dataset.data.median(numeric_only=True))
return dataset
@pipeline.add_node
def run_inference(dataset: Dataset) -> Dataset:
features = dataset.data[feature_names]
probabilities = model.predict_proba(features)[:, 1]
dataset.metadata["probabilities"] = probabilities
return dataset
return pipeline
How does FHIR become a DataFrame? A YAML schema maps FHIR resources to your training features:
# sepsis_vitals.yaml (excerpt)
features:
heart_rate:
fhir_resource: Observation
code: "220045" # MIMIC chartevents code
wbc:
fhir_resource: Observation
code: "51301" # MIMIC labevents code
age:
fhir_resource: Patient
field: birthDate
transform: calculate_age
No FHIR parsing code needed — define the mapping once, use it everywhere:
Explore Interactively
Step through the full flow in notebooks/fhir_ml_workflow.ipynb: FHIR bundle → Dataset → DataFrame → inference → RiskAssessment.
Bring your own model
The pre-generated model in cookbook/models/ is a synthetic demo — not trained on real patient data. To swap in your own:
import joblib
joblib.dump({
"model": your_trained_model, # any model with .predict_proba()
"metadata": {
"feature_names": ["heart_rate", "temperature", ...],
"metrics": {"optimal_threshold": 0.5}
}
}, "cookbook/models/sepsis_model.pkl")
Works with any scikit-learn-compatible model: XGBoost, LightGBM, or PyTorch/TF wrapped with a sklearn interface. To train on real MIMIC-IV data, see scripts/sepsis_prediction_training.py.
Pattern 1: Real-Time CDS Hooks Alerts
Use CDS Hooks when you need instant alerts during clinical workflows. The EHR triggers your service and pushes patient data via prefetch — no server queries needed.
Clinician opens chart → EHR fires patient-view hook → Your service runs prediction → CDS card appears in EHR
Set Up the CDS Hook Handler
Create a CDSHooksService that listens for patient-view events:
from healthchain.gateway import CDSHooksService
from healthchain.fhir import prefetch_to_bundle
from healthchain.models import CDSRequest, CDSResponse
from healthchain.models.responses.cdsresponse import Card
cds = CDSHooksService()
@cds.hook("patient-view", id="sepsis-risk")
def sepsis_alert(request: CDSRequest) -> CDSResponse:
if not request.prefetch:
return CDSResponse(cards=[])
# FHIR prefetch → Dataset → Prediction
bundle = prefetch_to_bundle(request.prefetch)
dataset = Dataset.from_fhir_bundle(bundle, schema=SCHEMA_PATH)
result = pipeline(dataset)
prob = float(result.metadata["probabilities"][0])
risk = "high" if prob > 0.7 else "moderate" if prob > 0.4 else "low"
if risk in ["high", "moderate"]:
return CDSResponse(cards=[
Card(
summary=f"Sepsis Risk: {risk.upper()} ({prob:.0%})",
indicator="critical" if risk == "high" else "warning",
detail=f"Predicted sepsis risk: {risk.upper()}. Recommend workup.",
source={"label": "HealthChain Sepsis Predictor"},
)
])
return CDSResponse(cards=[])
Build and Test the Service
Register with HealthChainAPI and test using the SandboxClient:
from healthchain.gateway import HealthChainAPI
app = HealthChainAPI(title="Sepsis CDS Hooks")
app.register_service(cds, path="/cds")
with app.sandbox("sepsis-risk") as client:
client.load_from_path("cookbook/data/mimic_demo_patients", pattern="*_patient.json")
responses = client.send_requests()
client.save_results("./output/")
Example CDS Response
{
"cards": [
{
"summary": "Sepsis Risk: HIGH (85%)",
"indicator": "critical",
"source": {
"label": "HealthChain Sepsis Predictor",
"url": "https://www.sccm.org/SurvivingSepsisCampaign/Guidelines/Adult-Patients"
},
"detail": "**AI Guidance:**\n- Predicted risk: **HIGH** (85%)\n- Recommend sepsis workup and early intervention.",
"title": "Sepsis Alert (AI Prediction)"
}
]
}
Advanced: Batch FHIR Gateway Screening
Use the FHIR Gateway when you need to screen multiple patients from a FHIR server. Unlike CDS Hooks (ephemeral alerts), this pattern persists predictions back to the FHIR server as RiskAssessment resources, making them available for dashboards, reports, and downstream workflows.
Prerequisites: A running FHIR server with patient data. This tutorial uses Medplum — see the FHIR Sandbox Setup guide to get credentials, then add them to .env:
MEDPLUM_BASE_URL=https://api.medplum.com/fhir/R4
MEDPLUM_CLIENT_ID=your_client_id
MEDPLUM_CLIENT_SECRET=your_client_secret
MEDPLUM_TOKEN_URL=https://api.medplum.com/oauth2/token
Upload demo patients to Medplum
Pre-extracted MIMIC demo patients are already in the repo. Upload them to your Medplum instance with:
The command prints the server-assigned IDs — copy them into DEMO_PATIENT_IDS in sepsis_fhir_batch.py:
✓ high_risk_bundle → PATIENT_ID=702e11e8-...
✓ low_risk_bundle → PATIENT_ID=3b0da7e9-...
✓ moderate_risk_bundle → PATIENT_ID=f490ceb4-...
To regenerate patients from a full MIMIC-on-FHIR dataset: python scripts/extract_mimic_demo_patients.py --minimal.
Screen Patients and Write Back Results
Configure the FHIRGateway, run predictions, and write RiskAssessment resources back to the server:
from healthchain.gateway import FHIRGateway
from healthchain.gateway.clients.fhir.base import FHIRAuthConfig
from healthchain.fhir.r4b import Patient, Observation
from healthchain.fhir import merge_bundles
gateway = FHIRGateway()
gateway.add_source("medplum", FHIRAuthConfig.from_env("MEDPLUM").to_connection_string())
def screen_patient(patient_id: str):
patient_bundle = gateway.search(Patient, {"_id": patient_id}, "medplum")
obs_bundle = gateway.search(Observation, {"patient": patient_id}, "medplum")
bundle = merge_bundles([patient_bundle, obs_bundle])
dataset = Dataset.from_fhir_bundle(bundle, schema=SCHEMA_PATH)
result = pipeline(dataset)
for ra in result.to_risk_assessment(
outcome_code="A41.9",
outcome_display="Sepsis",
model_name="sepsis_xgboost_v1",
):
gateway.create(ra, source="medplum")
for patient_id in DEMO_PATIENT_IDS:
screen_patient(patient_id)
Demo vs Production
This demo uses a fixed list of patient IDs. In production, query for patients dynamically — for example, ICU admissions in the last hour:
Expected Output
=== Screening patients from Medplum ===
702e11e8-...: HIGH (85%) → RiskAssessment/abc123
3b0da7e9-...: MODERATE (52%) → RiskAssessment/def456
f490ceb4-...: LOW (15%) → RiskAssessment/ghi789
RiskAssessment resources are visible in the Medplum console — search "RiskAssessment" in the resource type search bar.
Example RiskAssessment Resource
{
"resourceType": "RiskAssessment",
"id": "abc123",
"status": "final",
"subject": { "reference": "Patient/702e11e8-6d21-41dd-9b48-31715fdc0fb1" },
"method": {
"coding": [{
"system": "https://healthchain.io/models",
"code": "sepsis_xgboost_v1",
"display": "Sepsis XGBoost Model v1"
}]
},
"prediction": [{
"outcome": {
"coding": [{
"system": "http://hl7.org/fhir/sid/icd-10",
"code": "A41.9",
"display": "Sepsis"
}]
},
"probabilityDecimal": 0.85,
"qualitativeRisk": {
"coding": [{
"system": "http://terminology.hl7.org/CodeSystem/risk-probability",
"code": "high",
"display": "High likelihood"
}]
}
}]
}
What You've Built
Two deployment patterns for the same ML model:
| CDS Hooks | FHIR Gateway | |
|---|---|---|
| Integration | Event-driven (EHR pushes data) | Pull-based (service queries server) |
| Latency | Real-time (<1s) | Batch (seconds to minutes) |
| Output | CDS Cards (ephemeral alerts) | RiskAssessment (persisted resources) |
| Scaling | Per-patient on demand | Parallel/scheduled batch jobs |
Both patterns:
- Share the same model — train once, deploy multiple ways
- Use YAML feature schemas — declarative FHIR → features mapping, no custom parsing
- Handle FHIR natively — no custom data wrangling per integration
Use Cases
CDS Hooks (Real-time)
- Sepsis early warning alerts when opening ICU patient charts
- Drug interaction warnings during medication ordering
- Clinical guideline reminders triggered by diagnosis codes
FHIR Gateway (Batch)
- Nightly population health screening
- Quality measure calculation for reporting
- Research cohort identification
- Pre-visit risk stratification
Next Steps
- Bring your own model: Replace
sepsis_model.pklwith any scikit-learn-compatible model; updatesepsis_vitals.yamlto match your feature set - Add more FHIR sources: The gateway supports multiple sources — see the FHIR Sandbox Setup guide
- Combine patterns: Use batch screening to identify high-risk patients, then enable CDS alerts for those patients
- Automate batch runs: Schedule screening jobs with cron, Airflow, or cloud schedulers; or use FHIR Subscriptions to trigger on new ICU admissions (PRs welcome!)
- Go to production: Scaffold a project with
healthchain newand run withhealthchain serve— see From cookbook to service. Moving tohealthchain.yamlis where config-driven compliance support (audit logging, model versioning, deployment metadata) will live as those features mature.